RADseq and epiGBS identify population structure and DNA methylation along Coastal Louisiana
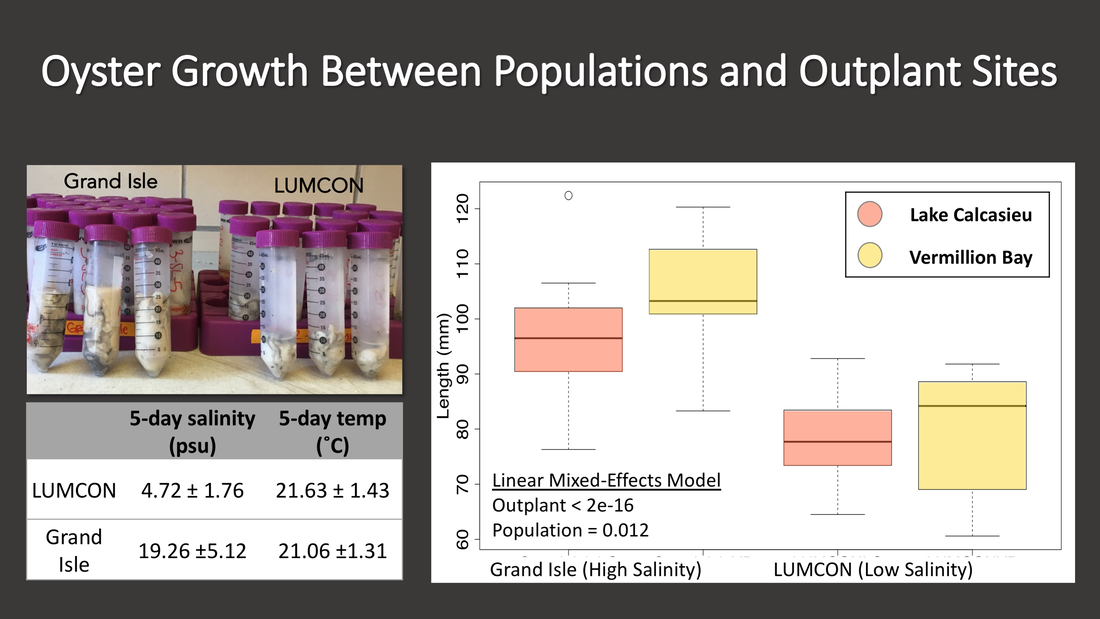
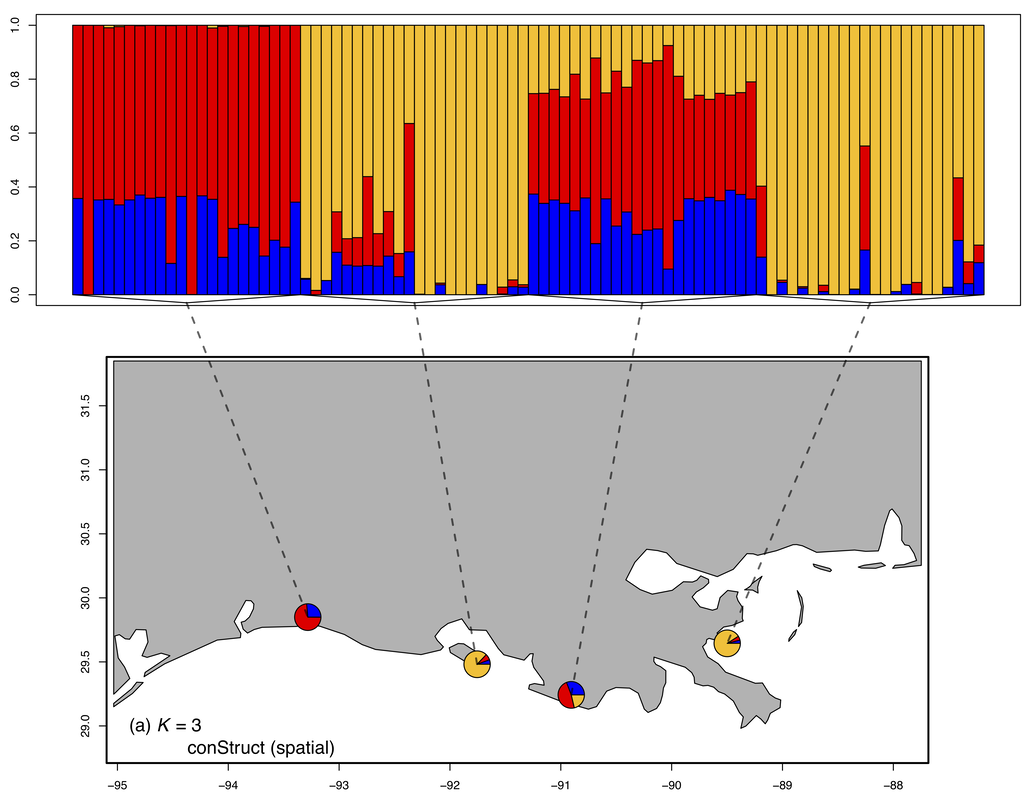
In this study we used a combination of RADseq and RRBS to examine the extent of genetic and epigenetic variation that was present between 4 populations of eastern oysters collected from across Louisiana. While we found evidence of population structure, the variation in DNA methylation was greater than the variation in SNPs. These results suggest that 5-10 % of DNA methylation is potentially environmentally plastic, while the remainder is genetically linked.
Johnson, KM, Kelly, MW. Population epigenetic divergence exceeds genetic divergence in the Eastern oyster Crassostrea virginica in the Northern Gulf of Mexico. Evol Appl. 2020; 00: 1– 15. https://doi.org/10.1111/eva.12912. [Open Access].
Salinity Responses to Temperature and Salinity in the eastern oyster.
In this study we used comparative transcriptomics and followed up these results with qPCR to measure the expression of a subset of genes of interest in wild caught individuals. These results highlight the negative impacts of increasing temperature and decreasing salinity on oyster reefs across Coastal Louisiana.
Jones, HR, Jones, KM, MW Kelly.Synergistic Effects of Temperature and Salinity on the Gene Expression and Physiology of Crassostrea virginica, Integrative and Comparative Biology, Volume 59, Issue 2, August 2019, Pages 306–319, https://doi.org/10.1093/icb/icz035
Comparative transcriptomics identifies signatures of temperature adaptation
In this study we have used comparative transcriptomics to identify change in gene expression between oysters from New Brunswick, Canada and Louisiana, USA. Despite inhabiting distinct thermal regimes, populations of C. virginica near the species’ southern and northern geographic range showed no physiological evidence of any population by temperature interactions. In this study, we used comparative transcriptomics to understand how oysters from either end of the species’ range maintain homeostasis across three acclimation temperatures (10, 20, and 30 ˚C). With this approach, we identified genes that were differentially expressed in response to temperature between individuals of C. virginica collected from New Brunswick, Canada and Louisiana, USA. We observed a larger core set of genes whose expression responded to temperature in both populations, and a smaller set of genes with expression patterns that were unique to each population. Intriguingly the genes with population-specific responses to temperature also had elevated FST values, but Ka/Ks ratios that were lower than the genome-wide average, suggesting that divergent FST may be due to selection in linked regulatory regions, rather than positive selection on protein coding sequences. Taken together our results suggest that, despite coarse-scale physiological similarities, natural selection on cis-regulatory regions has shaped divergent gene expression responses to temperature in geographically separated populations of this broadly eurythermal marine invertebrate.
Johnson, KM, Jones, HR, MW Kelly. Transcriptomic signatures of temperature adaptation in the eastern oyster Crassostrea virginica. In prep.
Epigenetic and transcriptomic signatures of Perkinsus marinus infection in the eastern oyster.
In this study we have used a combination of comparative epigentics and comparative trancsriptomics to examine the influence of dermo infection on the DNA methylation and transcritpomic phenotypes of oysters sourced from two of the distinct populations identified in our RADseq analysis.
Johnson, KM, Sirovy, KA, MW Kelly Characterizing the epigenetic and transcriptomic responses to Perkinsus marinus infection in the eastern oyster Crassostrea virginica. In prep
In this study we used a combination of RADseq and RRBS to examine the extent of genetic and epigenetic variation that was present between 4 populations of eastern oysters collected from across Louisiana. While we found evidence of population structure, the variation in DNA methylation was greater than the variation in SNPs. These results suggest that 5-10 % of DNA methylation is potentially environmentally plastic, while the remainder is genetically linked.
Johnson, KM, Kelly, MW. Population epigenetic divergence exceeds genetic divergence in the Eastern oyster Crassostrea virginica in the Northern Gulf of Mexico. Evol Appl. 2020; 00: 1– 15. https://doi.org/10.1111/eva.12912. [Open Access].
Salinity Responses to Temperature and Salinity in the eastern oyster.
In this study we used comparative transcriptomics and followed up these results with qPCR to measure the expression of a subset of genes of interest in wild caught individuals. These results highlight the negative impacts of increasing temperature and decreasing salinity on oyster reefs across Coastal Louisiana.
Jones, HR, Jones, KM, MW Kelly.Synergistic Effects of Temperature and Salinity on the Gene Expression and Physiology of Crassostrea virginica, Integrative and Comparative Biology, Volume 59, Issue 2, August 2019, Pages 306–319, https://doi.org/10.1093/icb/icz035
Comparative transcriptomics identifies signatures of temperature adaptation
In this study we have used comparative transcriptomics to identify change in gene expression between oysters from New Brunswick, Canada and Louisiana, USA. Despite inhabiting distinct thermal regimes, populations of C. virginica near the species’ southern and northern geographic range showed no physiological evidence of any population by temperature interactions. In this study, we used comparative transcriptomics to understand how oysters from either end of the species’ range maintain homeostasis across three acclimation temperatures (10, 20, and 30 ˚C). With this approach, we identified genes that were differentially expressed in response to temperature between individuals of C. virginica collected from New Brunswick, Canada and Louisiana, USA. We observed a larger core set of genes whose expression responded to temperature in both populations, and a smaller set of genes with expression patterns that were unique to each population. Intriguingly the genes with population-specific responses to temperature also had elevated FST values, but Ka/Ks ratios that were lower than the genome-wide average, suggesting that divergent FST may be due to selection in linked regulatory regions, rather than positive selection on protein coding sequences. Taken together our results suggest that, despite coarse-scale physiological similarities, natural selection on cis-regulatory regions has shaped divergent gene expression responses to temperature in geographically separated populations of this broadly eurythermal marine invertebrate.
Johnson, KM, Jones, HR, MW Kelly. Transcriptomic signatures of temperature adaptation in the eastern oyster Crassostrea virginica. In prep.
Epigenetic and transcriptomic signatures of Perkinsus marinus infection in the eastern oyster.
In this study we have used a combination of comparative epigentics and comparative trancsriptomics to examine the influence of dermo infection on the DNA methylation and transcritpomic phenotypes of oysters sourced from two of the distinct populations identified in our RADseq analysis.
Johnson, KM, Sirovy, KA, MW Kelly Characterizing the epigenetic and transcriptomic responses to Perkinsus marinus infection in the eastern oyster Crassostrea virginica. In prep